Super-resolution
The goal of this workflow aims at reconstructing high-resolution (HR) images from low-resolution (LR) ones.
Input:
LR image (single-channel or multi-channel). E.g. image with shape
(500, 500, 1)(y, x, channels)in2Dor(100, 500, 500, 1)(z, y, x, channels)in3D.
Output:
HR image.
In the figure below an example of this workflow’s input is depicted to make a x2 upsampling. The images were obtained from ZeroCostDL4Mic project:

Input LR image. |

Input HR image. |
Notice that the LR image has been resized but actually that is 502x502 whereas the HR is 1004x1004.
Data preparation
To ensure the proper operation of the library the data directory tree should be something like this:
Warning
Ensure that images and their corresponding masks are sorted in the same way. A common approach is to fill with zeros the image number added to the filenames (as in the example).
Configuration
Find in templates/super-resolution folder of BiaPy a few YAML configuration templates for this workflow.
Special workflow configuration
Here some special configuration options that can be selected in this workflow are described:
Upsampling is the most important variable to be set via
PROBLEM.SUPER_RESOLUTION.UPSCALING. In the example above, its value is2.Metrics: during the inference phase the performance of the test data is measured using different metrics if test masks were provided (i.e. ground truth) and, consequently,
DATA.TEST.LOAD_GTisTrue. In the case of super-resolution the Peak signal-to-noise ratio (PSNR) metrics is calculated when the HR image is reconstructed from individual patches.
Run
Select super-resolution workflow during the creation of a new configuration file:
Open a terminal as described in Installation. For instance, using 2d_super-resolution.yaml template file, the code can be run as follows:
# Configuration file
job_cfg_file=/home/user/2d_super-resolution.yaml
# Path to the data directory
data_dir=/home/user/data
# Where the experiment output directory should be created
result_dir=/home/user/exp_results
# Just a name for the job
job_name=my_2d_super_resolution
# Number that should be increased when one need to run the same job multiple times (reproducibility)
job_counter=1
# Number of the GPU to run the job in (according to 'nvidia-smi' command)
gpu_number=0
sudo docker run --rm \
--gpus "device=$gpu_number" \
--mount type=bind,source=$job_cfg_file,target=$job_cfg_file \
--mount type=bind,source=$result_dir,target=$result_dir \
--mount type=bind,source=$data_dir,target=$data_dir \
BiaPyX/biapy \
-cfg $job_cfg_file \
-rdir $result_dir \
-name $job_name \
-rid $job_counter \
-gpu $gpu_number
Note
Note that data_dir must contain all the paths DATA.*.PATH and DATA.*.GT_PATH so the container can find them. For instance, if you want to only train in this example DATA.TRAIN.PATH and DATA.TRAIN.GT_PATH could be /home/user/data/train/x and /home/user/data/train/y respectively.
Open a terminal as described in Installation. For instance, using 2d_super-resolution.yaml template file, the code can be run as follows:
# Configuration file
job_cfg_file=/home/user/2d_super-resolution.yaml
# Where the experiment output directory should be created
result_dir=/home/user/exp_results
# Just a name for the job
job_name=my_2d_super_resolution
# Number that should be increased when one need to run the same job multiple times (reproducibility)
job_counter=1
# Number of the GPU to run the job in (according to 'nvidia-smi' command)
gpu_number=0
# Move where BiaPy installation resides
cd BiaPy
# Load the environment
conda activate BiaPy_env
python -u main.py \
--config $job_cfg_file \
--result_dir $result_dir \
--name $job_name \
--run_id $job_counter \
--gpu $gpu_number
For multi-GPU training you can call BiaPy as follows:
gpu_number="0, 1, 2"
python -u -m torch.distributed.run \
--nproc_per_node=3 \
main.py \
--config $job_cfg_file \
--result_dir $result_dir \
--name $job_name \
--run_id $job_counter \
--gpu $gpu_number
nproc_per_node need to be equal to the number of GPUs you are using (e.g. gpu_number length).
Results
The results are placed in results folder under --result_dir directory with the --name given. An example of this workflow is depicted below:
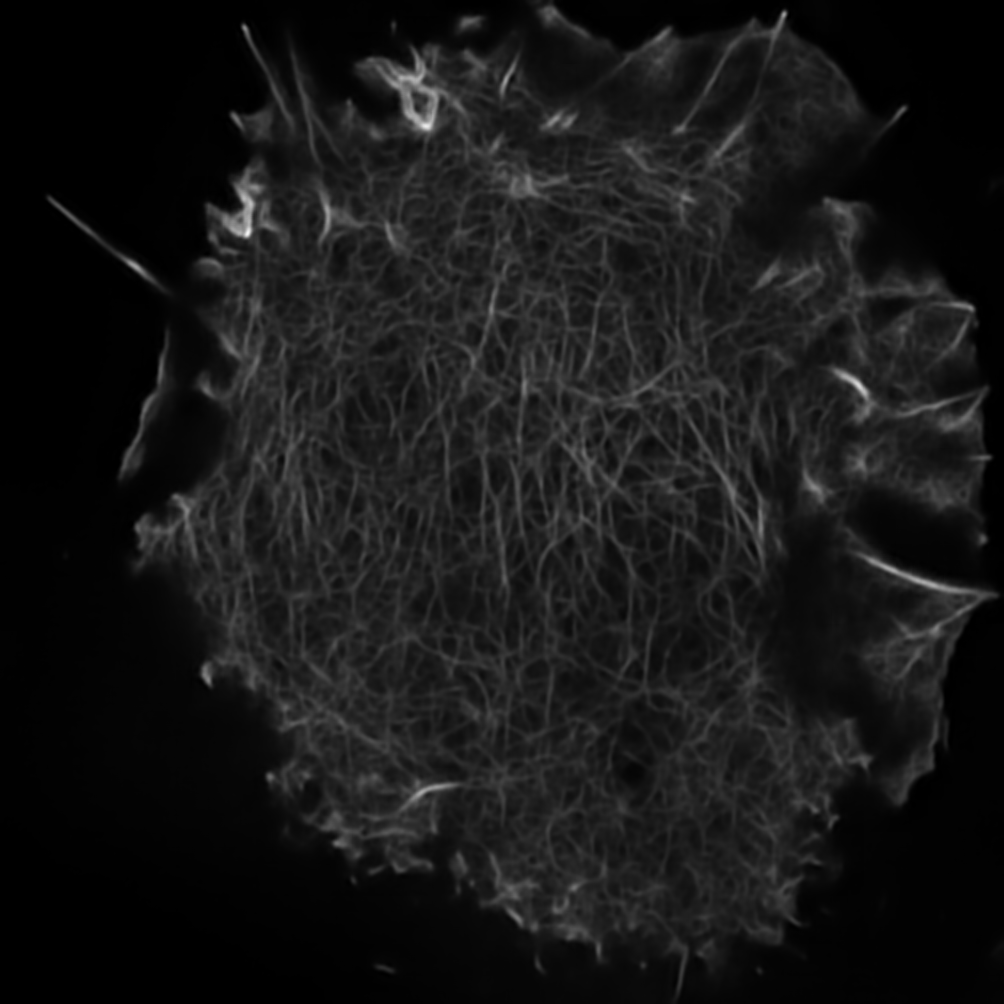

Predicted HR image. |

Input HR image. |
Here both images are of size 1004x1004.
Following the example, you should see that the directory /home/user/exp_results/my_2d_super_resolution has been created. If the same experiment is run 5 times, varying --run_id argument only, you should find the following directory tree:
config_files: directory where the .yaml filed used in the experiment is stored.2d_super-resolution.yaml: YAML configuration file used (it will be overwrited every time the code is run)
checkpoints: directory where model’s weights are stored.my_2d_super-resolution_1-checkpoint-best.pth: checkpoint file (best in validation) where the model’s weights are stored among other information.normalization_mean_value.npy: normalization mean value (only created ifDATA.NORMALIZATION.TYPEiscustom). Is saved to not calculate it everytime and to use it in inference.normalization_std_value.npy: normalization std value (only created ifDATA.NORMALIZATION.TYPEiscustom). Is saved to not calculate it everytime and to use it in inference.
results: directory where all the generated checks and results will be stored. There, one folder per each run are going to be placed.my_2d_super_resolution_1: run 1 experiment folder.aug: image augmentation samples.charts:my_2d_super_resolution_1_*.png: Plot of each metric used during training.my_2d_super_resolution_1_loss.png: Loss over epochs plot (when training is done).model_plot_my_2d_super_resolution_1.png: plot of the model.
per_image:.tif files: reconstructed images from patches.
train_logs: each row represents a summary of each epoch stats. Only avaialable if training was done.tensorboard: Tensorboard logs.
Note
Here, for visualization purposes, only my_2d_super_resolution_1 has been described but my_2d_super_resolution_2, my_2d_super_resolution_3, my_2d_super_resolution_4 and my_2d_super_resolution_5 will follow the same structure.